Parameters
Parameter Name | Variable | Default Value | Parameter Range | Description |
|---|---|---|---|---|
NB_MOLECULES | 5,000,000 | >0 | number of expressed RNA molecules simulated | |
EXPRESSION_K | -0.6 | exponent of the expression power law ("Pareto coefficient") | ||
EXPRESSION_X0 |
| 9,500 | parameter of the exponential decay | |
EXPRESSION_X1 | 9,5002 | parameter of the exponential decay |
Algorithm
Input: reference annotation (REF_FILE), transcript filtering parameter (LOAD_CODING, LOAD_NONCODING), expression parameters (NB_MOLECULES, EXPRESSION_K, EXPRESSION_X0, EXPRESSION_X1)
In the beginning, the Flux Simulator reads the transcripts of the reference annotation (REF_FILE) and clusters genomic overlapping ones into loci. Transcripts that are annotated as non-/coding can be selectively disregarded (LOAD_CODING, LOAD_NONCODING). Then to assign a random expression profile where not necessarily all transcripts of the reference are expressed. Expression levels are connected with the relative expression rank
by a mixed power- and exponential law of the general form
where denotes the rank number of a gene,
is the exponent of the intrinsic power law, and
respectively
control the exponential decay. The Flux Simulator assigns to the transcripts in the reference annotation randomly expression ranks
which then are turned into relative expression levels by the modified Zipf's Law above, which determines the initial number of molecules by multiplication with the total numbers of molecules. Default values for parameters
and
have been estimated for mammalian cells by non-linear fitting to expression levels observed in experimental results.
Output: Columnn 1-6 of the PRO_FILE, i.e., (1) locus name, (2) transcript identifier, (3) coding flag, (4) length of the processed transcript, (5) relative fraction and (6) absolute number of the transcript species in the initial RNA extraction.
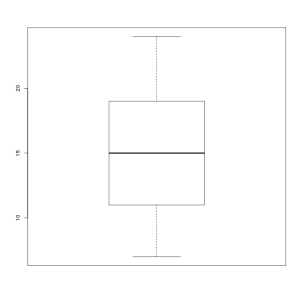
| Although the Simulator program currently allows to set parameter EXPRESSION_K to a value of "0", please note that such settings remove the power-law distribution we ususally observe in cellular transcriptomes. Below a boxplot representation of the distribution of the simulated number of molecules for all transcripts in a reference annotation when EXPRESSION_K is set to "0", the distribution exhibits a mean of ~15 and a standard deviation of ~5. |